
Figure 1The distribution of the number of viruses detected per participant (doughnut chart) and the number of different virus species detected overall (bar chart) is shown for (A) all viruses and (B) viruses causing respiratory tract infections.
DOI: https://doi.org/https://doi.org/10.57187/s.3797
Acute respiratory tract infections are among the most common reasons for consultations in the outpatient setting [1] and the leading cause of antibiotic prescription and use [2–4]. While patients with pneumonia and chronic obstructive pulmonary disease (COPD) exacerbations benefit from antibiotic treatment, most respiratory tract infections will resolve spontaneously [3] since they are predominantly caused by viruses [4]. Rapid and comprehensive diagnostic testing, informing healthcare workers about the suspected cause of disease, may reduce diagnostic uncertainty. Currently, acute respiratory tract infections are diagnosed predominantly based on clinical assessment. Microbial studies other than influenza-specific assays or rapid antigen detection for Streptococcus pyogenes are not routinely performed. Consequently, the causal pathogen is rarely identified. Therefore, timely pan-pathogen detection could greatly help direct the treatment choice.
Metagenomic next-generation sequencing (mNGS) allows for the comprehensive analysis of microbial material from a clinical specimen. Unlike conventional diagnostic tests, which target specific pathogens and rely on a working hypothesis, mNGS enables the detection of unexpected or even unknown pathogens [5]. The utility of mNGS in identifying infectious agents has previously been demonstrated for diseases such as meningitis and encephalitis [6–8], respiratory infections [9, 10], joint infection [11], and endocarditis [12]. Falling sequencing costs and methods adapted for diagnostic use have now made it financially feasible for routine testing. Nonetheless, mNGS is mainly used in difficult-to-diagnose cases in an inpatient setting.
Here, we evaluated the potential of viral mNGS as a diagnostic tool in primary care for immunocompetent patients suffering from a respiratory tract infection.
This prospective cross-sectional study included patients presenting to their general practitioner with symptoms of a respiratory tract infection from October 2019 to December 2020. Its primary outcome was the identification of viral pathogens from throat swabs by viral mNGS. Its secondary outcomes were the frequency of viral pathogen detection and the correlation between the routine clinical diagnosis and the detected viral pathogens. Written informed consent was obtained from all study participants. The study protocol was approved by the Ethics Committee of the canton of Zurich (BASEC-Nr. 2019-01120, 19.08.2018).
Twenty-one primary care physicians from the cantons of Zurich and Basel-Landschaft were selected from physicians known to the authors based on their interest in the study and willingness to enrol patients in a busy environment. Patients aged ≥18 years with clinically suspected respiratory tract infections were eligible for enrollment. Exclusion criteria were limited German language skills, pregnancy, previous enrollment in the trial, and immunosuppression (treatment with biologicals, cytostatic drugs, prednisone >20 mg for >4 weeks, or glucocorticoid therapy at equivalent dosage). Patients within practices were randomly selected based on their presentation time and according to the inclusion/exclusion criteria.
During the consultation, the primary care physician or a nurse collected throat swabs (FLOQSwabs with UTM-RT transport medium for viruses; Copan, Italy) from all study participants. The physician also completed a questionnaire gathering demographic and medical data, including information about the suspected disease aetiology and planned treatment. Routine care was provided at the general practitioner’s discretion.
On day 14 after the initial consultation, a trained study physician collected information on the clinical course through a structured telephone interview with the participant. Unavailable patients were repeatedly called with at least four attempts.
Throat swabs were analysed at the Institute of Medical Virology (Zurich, Switzerland) using a viral mNGS approach, as previously described [8] (https://github.com/medvir/virome-protocols/releases/tag/v2.2.1). Briefly, samples were pre-processed upon arrival by centrifugation and filtration (0.45 µm) and stored at –80 °C. Total nucleic acids were extracted using the NucliSENS EasyMAG System (BioMérieux, Craponne, France). Next, reverse transcription with random hexamers and second-strand synthesis were performed separately for the RNA and DNA genomes. Then, sequencing libraries were constructed using the NexteraXT protocol (Illumina, San Diego, CA, USA) and sequenced on an Illumina MiSeq system for 1 × 151 single-end cycles using version 3 chemistry. A maximum of five samples (plus a negative control) were sequenced per run. Reads were analysed with a dedicated bioinformatic pipeline called VirMet (github.com/medvir/VirMet/releases/tag/v1.1.1). Samples sequenced after June 2020 included an internal RNA positive control. All runs included a negative control. Viral sequence data has been uploaded to Zenodo (doi 10.5281/zenodo.7185259).
The VirMet output for each sequencing run was evaluated for total and quality filtered reads per sample, the distribution of reads into different taxonomic categories (viral, bacterial, human, and unknown origin), and viruses detected at the species level and corresponding reference sequences. Positive virus hits were based on the number of reads with a threshold of at least three reads per species, distribution of the reads across the viral genome, and detection in the corresponding workflow (DNA/RNA), as previously described [8]. Final sequencing results were made available to the general practitioner retrospectively.
The respiratory viruses detected by mNGS were validated using a conventional syndromic testing panel and a SARS-CoV-2 polymerase chain reaction (PCR) assay. The FDA and CE-IVD-approved ePlex Respiratory Pathogen Panel (GenMark Diagnostics, Carlsbad, CA, USA) can detect 25 respiratory pathogens using an electrochemical detection system. Viral targets include human adenovirus, human coronaviruses 299E/HKU1//NL63/OC43, Middle East respiratory syndrome coronavirus, human bocavirus, human metapneumovirus, human rhinovirus/enterovirus, influenza A virus, influenza B virus, parainfluenza viruses 1/2/3/4, and respiratory syncytial viruses A/B. The tests were performed according to the manufacturer’s instructions on all available throat swabs with archived material (n = 275). All available samples collected after January 2020 (n = 111) were validated for SARS-CoV-2 using the Cobas SARS-CoV-2 IVD test (Roche Diagnostics GmbH) on a Cobas 6800, as previously described [13]. The performance of mNGS was evaluated by calculating the positive and negative percentage agreement (PPA/NPA).
Discrepant results between mNGS and ePlex were confirmed using the five-tube multiplex FTD Respiratory Pathogens 21 RT-PCR assay run on a ViiA7 Real-Time PCR System (Life Technologies, Carlsbad, CA, USA) using AgPath-IDTM One-Step RT-PCR Reagents (Ambion, Thermo Fisher Scientific, Waltham, MA, USA). The following targets of the FTD21-assay were included in the analysis: human adenovirus, human coronavirus HKU1, human rhinovirus, human enterovirus, influenza A virus, and parainfluenza viruses 2/4. For sensitivity reasons, an additional RT-PCR to detect human rhinovirus was included in the multiplex assay, as described previously [14].
Descriptive summaries are reported as counts and percentages for categorical variables and median and interquartile range (IQR) for continuous variables. The Shapiro-Wilk test for normality was performed using the R statistical software (version 4.0.2).
Two hundred eighty patients were enrolled from October 2019 to December 2020. Three were subsequently excluded (one due to pregnancy and two due to age <18 years). The remaining 277 patients provided throat swabs, which were analysed by mNGS. Two hundred sixty-four patients (95%) completed a follow-up on the clinical course. Slightly more women (n = 162, 58.5%) were enrolled (table 1). The participants’ median age was 39 years (IQR = 30–52). Ninety-five participants (34%) had one or more known comorbidities. Only 35 participants (12.6%) had been vaccinated against influenza in the respective season. At the first outpatient visit, 188 participants (68%) reported having a “sore throat”, which was the most frequently reported symptom. Other common clinical presentations were, in decreasing order of frequency, rhinitis, dry cough, headache, and myalgia. Only 17 of the 241 participants (7%) with recorded body temperature presented with a temperature above 38 °C. The participants’ median duration of respiratory tract infection symptoms before the first outpatient visit was five days. The most common postulated clinical diagnoses were common cold/rhinitis, non-streptococcal pharyngitis/tonsillitis, and acute bronchitis (table 1).
Table 1Participants’ demographics and baseline clinical findings.
| Baseline characteristics | Total number of participants, n | 277 | |
| Age, in years, median (IQR) | 39 (30–52) | ||
| Female, n (%) | 162 (58.5%) | ||
| Comorbidities, n (%) | Total patients with known comorbidities | 95 (34.3%) | |
| Cardiovascular disease | 27 (28.4%) | ||
| Pulmonary disease | 23 (24.2%) | ||
| Other diseases | 45 (47.4%) | ||
| No known comorbidities | 92 (33.6%) | ||
| Unknown | 90 (32.1%) | ||
| Vaccination status, n (%) | Influenza vaccinated | 35 (12.6%) | |
| Symptoms/clinical findings | Symptom duration, in days, median (IQR) | 5 (3–7) | |
| Temperature, median (IQR) | 37 (36.6–37.4) | ||
| Sore throat, n (%) | 188 (67.9%) | ||
| Rhinitis, n (%) | 163 (58.8%) | ||
| Dry cough, n (%) | 136 (49.1%) | ||
| Headache, n (%) | 133 (48.0%) | ||
| Myalgia, n (%) | 120 (43.3%) | ||
| Fever, n (%) | 88 (31.8%) | ||
| Productive cough, n (%) | 74 (26.7%) | ||
| Dysphagia, n (%) | 42 (15.2%) | ||
| Ear pain, n (%) | 34 (12.3%) | ||
| Dyspnoea, n (%) | 25 (9.0%) | ||
| Other, n (%) | 40 (14.4%) | ||
| Presumed clinical diagnosis, n (%) | Common cold/rhinitis | 136 (49.1%) | |
| Pharyngitis/tonsillitis (non-streptococcal) | 44 (15.9%) | ||
| Acute bronchitis | 31 (11.2%) | ||
| Influenza | 22 (7.9%) | ||
| COVID-19 | 15 (5.4%) | ||
| Streptococcal pharyngitis | 6 (2.2%) | ||
| Acute sinusitis | 5 (1.8%) | ||
| Community-acquired pneumonia | 5 (1.8%) | ||
| Exacerbated asthma | 2 (0.7%) | ||
| Exacerbated COPD | 2 (0.7%) | ||
| Other | 9 (3.2%) | ||
| Unknown | 26 (9.4%) | ||
COPD: chronic obstructive pulmonary disease; COVID-19: coronavirus disease 2019; IQR: interquartile range.
Laboratory workup was performed for 153 participants (55.2%). Out of 148 C-reactive protein (CRP) measurements, 99 (66.9%) were elevated above a threshold of 5 mg/l. Leukocytosis (defined as leukocyte count exceeding 10 G/l according to Schernberg et al. [15]) was rarely observed, with 21 cases (14.3%) out of 147 measurements (data not shown).
Viral mNGS of throat swabs was performed for all 277 participants. DNA and RNA sequencing reads per sample and reads per detected virus are provided in table S1 in the Appendix. At least one virus was detected for 206 participants (74%). mNGS detected 357 viral species overall (figure 1A), with a maximum of six per participant. We found that 164 (46%) virus sequences belonged to the herpesviridae family, of which 120 (73%) were identified as human betaherpesvirus 7 (HHV-7). Most importantly, 145 (40.6%) of the sequences referred to viruses considered causes of respiratory tract infections, detected in 138 participants (49.8%).

Figure 1The distribution of the number of viruses detected per participant (doughnut chart) and the number of different virus species detected overall (bar chart) is shown for (A) all viruses and (B) viruses causing respiratory tract infections.
In the next step, we only analysed viruses likely to cause respiratory tract infections, which will be referred to as “respiratory viruses” hereafter. We detected one respiratory virus in 131 samples (47.3% of participants) and more than one in only seven samples (2.5% of participants). Rhinoviruses (including rhinovirus A, B, and C) were the most frequently identified respiratory viruses, accounting for 56 sequences and 38.6% of all 145 viruses detected as potentially causing respiratory tract infections (20.2% of participants; figure 1B). Overall, most virus sequences were detected in the winter months of 2019/2020 (figure 2). For example, the influenza virus detections were confined to December to March, correlating with the seasonal pattern of the influenza epidemic in Switzerland [16]. From March 2020, we observed an overall reduction in patient enrollment (figure S1A in the Appendix) and an initial reduction in respiratory virus detections (figure S1B in the Appendix), coinciding with the first SARS-CoV-2 cases. The mean proportion of positive study visits over the entire study period was 46%, with a minimum of 11% in May 2020 (figure S1B in the Appendix). All SARS-CoV-2 detections clustered to the time of the first and second waves of the pandemic in Switzerland in the spring and fall of 2020. Notably, while rhinovirus sequences were detected over the entire study period, the detection of the other respiratory viruses declined after March 2020, including influenza A virus (figure 2).
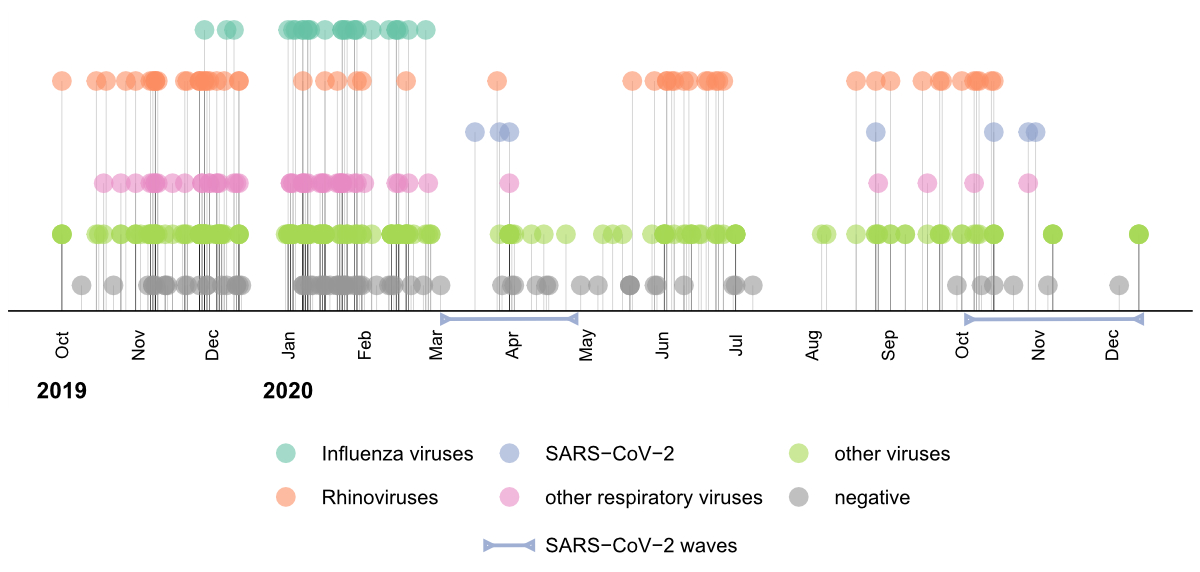
Figure 2A timeline of the study visits and detected viruses throughout the study period from October 2019 to December 2020. Negative and positive sequencing results (grey and coloured circles, respectively) are shown for each study visit. The detection of influenza viruses (green), rhinoviruses (orange), SARS-CoV-2 (blue), respiratory viruses other than rhinoviruses and SARS-CoV-2 (pink), and other non-respiratory viruses (green) is highlighted separately. The two SARS-CoV-2 waves in Switzerland within the studied period are indicated by blue bars.
Out of all respiratory tract infections (n = 277), the general practitioners suspected a viral cause in 252 cases (91%), whereas 21 (7.6%) were believed to be of bacterial origin, one (0.4%) was thought to have another cause, and three (1%) were not specified. mNGS detected a respiratory virus in 133 participants (52.8%) with an initially postulated viral cause (n = 252), and 119 participants (47.2%) remained negative for respiratory viruses (table 2). For participants where the general practitioner suspected a bacterial cause (n = 21), mNGS detected a respiratory virus in five (23.8%), and 16 (76.2%) remained negative for respiratory viruses. The presumed clinical diagnoses were acute bronchitis in two of the mNGS-positive participants with detection of influenza A virus and rhinovirus A, pneumonia in one participant with detection of human metapneumovirus, streptococcal pharyngitis in one participant with detection of enterovirus B, and tonsillitis in one participant with detection of human mastadenovirus C (table S2 in the Appendix). Overall, 138 infections (49.8%) could have been attributed to a viral cause based on mNGS, and antibiotic prescription could have been avoided in five (20.8%) of the 24 participants with a suspected bacterial cause who were prescribed antibiotics.
Table 2Postulated cause of respiratory infection and mNGS results for respiratory viruses.
| Postulated cause | n (%) | mNGS positive | mNGS negative |
| Viral | 252 (91.0%) | 133 | 119 |
| Bacterial | 21 (7.6%) | 5 | 16 |
| Other/not specified | 4 (1.4%) | 0 | 4 |
mNGS: metagenomic next-generation sequencing
We reanalysed all available samples using the ePlex Respiratory Pathogen Panel and the Cobas SARS-CoV-2 IVD assay to assess the performance of mNGS compared to conventional testing for detecting respiratory pathogens. For the 275 samples and 17 viral targets tested, PPA (sensitivity) was 89.1%, and NPA (specificity) was 99.8% (table 3). To disentangle differences in performance regarding specific viral targets, we calculated PPA and NPA for the most frequently detected viruses in the tested panel: human rhinovirus/enterovirus, influenza A virus, human coronavirus HKU1, and SARS-CoV-2. PPA was lowest for the influenza A virus at 83.3%, and NPA was lowest for the human rhinovirus/enterovirus at 98.6%. Overall, the percentage agreement was at least 97.8% for all four targets (table 3).
Table 3Performance comparison between mNGS and conventional testing for respiratory infections (ePlex and SARS-CoV-2 IVD test) for all targets and the most frequently detected targets in the panel.
| Conventional test (ePlex, Cobas) | |||||
| + | – | ||||
| All targets | OPA = 99.5% | PPA = 89.1% | mNGS + | 115 | 9 |
| NPA = 99.8% | mNGS – | 14 | 4373 | ||
| Rhino/enterovirus | OPA = 97.8% | PPA = 94.8% | mNGS + | 55 | 3 |
| NPA = 98.6% | mNGS – | 3 | 214 | ||
| Influenza A | OPA = 98.2% | PPA = 83.3% | mNGS + | 20 | 1 |
| NPA = 99.6% | mNGS – | 4 | 250 | ||
| huCoV HKU1 | OPA = 99.3% | PPA = 88.9% | mNGS + | 8 | 1 |
| NPA = 99.6% | mNGS – | 1 | 265 | ||
| SARS-CoV-2 | OPA = 99.1% | PPA = 87.5% | mNGS + | 7 | 0 |
| NPA = 100% | mNGS – | 1 | 103 | ||
huCoV HKU1: human coronavirus HKU1; mNGS: metagenomic next-generation sequencing; NPA: negative percentage agreement; OPA: overall percentage agreement; PPA: positive percentage agreement; SARS-CoV-2: severe acute respiratory syndrome-related coronavirus type 2.
In total, 23 discrepancies were observed, which were resolved using specific PCR for the respective viruses as the reference standard. Of the 14 samples that tested positive by ePlex but not by mNGS, six could not be confirmed by specific PCR and were rated as false positive by ePlex, five had respective viral reads detected by mNGS but below our internal threshold of at least three reads, and three had low total mNGS read counts in the affected RNA sample (<15,000). Of nine samples that tested positive by mNGS but not by ePlex, eight could be confirmed by specific PCR and were rated as false negative by ePlex, and one was false positive with mNGS. A low initial viral load or nucleic acid degradation before retesting on the ePlex might explain this presumably false positive result for this sample.
In this study, we investigated the application of mNGS for identifying viral pathogens in outpatients with upper respiratory tract infections. Half of the suspected respiratory tract infections in the studied patients could be ascribed to a viral origin through mNGS. The identification of numerous respiratory virus species underscores the comprehensive character of the metagenomic approach and its potential to directly guide upper respiratory tract infection treatment and prevent unnecessary antibiotic prescription. Moreover, we were able to highlight the seasonality of viruses circulating in the community, as demonstrated by the examples of influenza and other respiratory viruses. As highlighted by our study’s early detection of SARS-CoV-2, mNGS is beneficial for identifying novel pathogens. Indeed, SARS-CoV-2 was identified using mNGS [17]. Further into the pandemic, we likely underestimate the number of SARS-CoV-2 infections due to the many easily accessible testing opportunities.
Both early identification and surveillance of respiratory viruses are essential to guide disease mitigation and public health interventions [18, 19]. The comprehensive pathogen detection by mNGS could provide further insight into the effect and implementation of non-pharmaceutical hygienic measures. Despite interventions such as social distancing, hand hygiene, and mandatory use of facemasks implemented by the Swiss government [20, 21], continuous circulation of rhinoviruses was observed.
Notably, while mask policies were implemented later in Switzerland than elsewhere, the lifting of COVID-19 non-pharmaceutical interventions led to an inter-seasonal surge in respiratory syncytial virus (RSV) infection internationally and in Switzerland in 2021 [22]. The comprehensive character of mNGS, while not planned but still demonstrated in our study, would be ideal to rapidly detect such upcoming epidemics when used in a sentinel surveillance approach for respiratory and emerging viruses in the community, allowing the detection of novel or even unknown pathogens.
The performance comparison of mNGS and conventional testing showed high agreement, with an overall PPA of 89.1%. Discrepant results were primarily due to decreased sensitivity in either platform. However, the advantage of mNGS over syndromic panels is the inclusion of an unlimited number of pathogens, the detection of potential novel viruses, and even the definition of variants and subtypes if coverage is sufficient.
Overall, most detected sequences were for viruses not known to cause respiratory tract infections. The abundance of herpesvirus sequences, with a clear dominance of HHV-7, is a consequence of herpesvirus latency. The salivary glands have been proposed to be the anatomic location of HHV-7 latency. Continuous shedding of HHV-7 in healthy adults has been reported, with a frequency of about 34% [23, 24], comparable to our results. These findings also show that the clinical significance of pathogens detected by mNGS must be carefully interpreted.
In this study, we provided the clinician and patient with retrospective information on detected virus sequences and potential causes of the respiratory tract infection. Today, mNGS remains a time-consuming and complex method requiring benchtop sequencers and bioinformatic analysis. Technological advances are needed to reduce the turnaround time, complexity, and costs of mNGS to provide timely and actionable results. These advances are within reach with real-time data from Nanopore sequencing, which provides results in as little as six hours [25, 26]. While further improvements are needed for mNGS to fully develop as a primary diagnostic tool at the patient level, its retrospective analysis and surveillance capabilities are already insurmountable and must be leveraged for epidemiologic surveillance.
We thank all the participating patients and general practitioners: Dr. med. Ute Plüss, Dr. med. Monika Witzig, Dr. med. Anitha Vilan Bossi, Dr. med. Andreas Schindler, Dr. med. Verena Meyboom, Dr. med. Anita Fröhlich, Dr. med. Simon Brüllmann, Dr. med. Claudia Langer, prakt med Olivier Diener, Dr. med. Katja Bauder, Dr. med. Matthias Günthard, Dr. med. Hans-Martin Maurer, Dr. med. Regula Capaul, Dr. med. Roman Kind, Dr. med. Anne Brausch, Dr. med. Anja Weibel, Dr. med. Peter Ramer, Dr. med. Andreas Steiner, Dr. med. Anna Bettina Hunger, Dr. med. Silvia Meierhans, and Dr. med. Rahel Hottinger.
This work was supported by the Clinical Research Priority Program “Comprehensive Genomic Pathogen Detection” of the University of Zurich.
All authors have completed and submitted the International Committee of Medical Journal Editors form for disclosure of potential conflicts of interest. No potential conflict of interest related to the content of this manuscript was disclosed.
1. Finley CR, Chan DS, Garrison S, Korownyk C, Kolber MR, Campbell S, et al. What are the most common conditions in primary care? Systematic review. Can Fam Physician. 2018 Nov;64(11):832–40.
2. Gulliford MC, Dregan A, Moore MV, Ashworth M, Staa TV, McCann G, et al. Continued high rates of antibiotic prescribing to adults with respiratory tract infection: survey of 568 UK general practices. BMJ Open. 2014 Oct;4(10):e006245.
3. Thompson W, Tonkin-Crine S, Pavitt SH, McEachan RR, Douglas GV, Aggarwal VR, et al. Factors associated with antibiotic prescribing for adults with acute conditions: an umbrella review across primary care and a systematic review focusing on primary dental care. J Antimicrob Chemother. 2019 Aug;74(8):2139–52.
4. Shaver AL, Jacobs DM, LaMonte MJ, Noyes K. Antibiotic prescribing for acute respiratory tract infections in the United States outpatient setting. BMC Fam Pract. 2019 Jul;20(1):91.
5. Chiu CY, Miller SA. Clinical metagenomics. Nat Rev Genet. 2019 Jun;20(6):341–55.
6. Wilson MR, Sample HA, Zorn KC, Arevalo S, Yu G, Neuhaus J, et al. Clinical Metagenomic Sequencing for Diagnosis of Meningitis and Encephalitis. N Engl J Med. 2019 Jun;380(24):2327–40. 10.1056/NEJMoa1803396
7. Tschumi F, Schmutz S, Kufner V, Heider M, Pigny F, Schreiner B, et al. Meningitis and epididymitis caused by Toscana virus infection imported to Switzerland diagnosed by metagenomic sequencing: a case report. BMC Infect Dis. 2019 Jul;19(1):591.
8. Kufner V, Plate A, Schmutz S, Braun DL, Günthard HF, Capaul R, et al. Two Years of Viral Metagenomics in a Tertiary Diagnostics Unit: Evaluation of the First 105 Cases. Genes (Basel). 2019 Aug;10(9):661.
9. Langelier C, Kalantar KL, Moazed F, Wilson MR, Crawford ED, Deiss T, et al. Integrating host response and unbiased microbe detection for lower respiratory tract infection diagnosis in critically ill adults. Proc Natl Acad Sci USA. 2018 Dec;115(52):E12353–62.
10. Zinter MS, Dvorak CC, Mayday MY, Iwanaga K, Ly NP, McGarry ME, et al. Pulmonary Metagenomic Sequencing Suggests Missed Infections in Immunocompromised Children. Clin Infect Dis. 2019 May;68(11):1847–55.
11. Thoendel MJ, Jeraldo PR, Greenwood-Quaintance KE, Yao JZ, Chia N, Hanssen AD, et al. Identification of Prosthetic Joint Infection Pathogens Using a Shotgun Metagenomics Approach. Clin Infect Dis. 2018 Oct;67(9):1333–8.
12. Million M, Gaudin M, Melenotte C, Chasson L, Edouard S, Verdonk C, et al. Metagenomic Analysis of Microdissected Valvular Tissue for Etiological Diagnosis of Blood Culture-Negative Endocarditis. Clin Infect Dis. 2020 May;70(11):2405–12.
13. Huber M, Schreiber PW, Scheier T, Audigé A, Buonomano R, Rudiger A, et al. High Efficacy of Saliva in Detecting SARS-CoV-2 by RT-PCR in Adults and Children. Microorganisms. 2021 Mar;9(3):642.
14. Steiner F, Schmutz S, Gosert R, Huder JB, Redli PM, Capaul R, et al. Usefulness of the GenMark ePlex RPP assay for the detection of respiratory viruses compared to the FTD21 multiplex RT-PCR. Diagn Microbiol Infect Dis. 2021 Sep;101(1):115424.
15. Schernberg A, Blanchard P, Chargari C, Ou D, Levy A, Gorphe P, et al. Leukocytosis, prognosis biomarker in locally advanced head and neck cancer patients after chemoradiotherapy. Clin Transl Radiat Oncol. 2018 Jul;12:8–15.
16. Saisonale Grippe - Lagebericht Schweiz. Available: https://www.bag.admin.ch/bag/de/home/krankheiten/ausbrueche-epidemien-pandemien/aktuelle-ausbrueche-epidemien/saisonale-grippe---lagebericht-schweiz.html
17. Wu F, Zhao S, Yu B, Chen YM, Wang W, Song ZG, et al. A new coronavirus associated with human respiratory disease in China. Nature. 2020 Mar;579(7798):265–9.
18. Fox JD. Respiratory virus surveillance and outbreak investigation. J Clin Virol. 2007 Nov;40 Suppl 1:S24–30. 10.1016/S1386-6532(07)70006-9
19. Heeney JL. Zoonotic viral diseases and the frontier of early diagnosis, control and prevention. J Intern Med. 2006 Nov;260(5):399–408.
20. Nadeau SA, Vaughan TG, Beckmann C, Topolsky I, Chen C, Hodcroft E, et al. Swiss public health measures associated with reduced SARS-CoV-2 transmission using genome data. Sci Transl Med. 2023 Jan;15(680):eabn7979.
21. Plümper T, Neumayer E. Lockdown policies and the dynamics of the first wave of the Sars-CoV-2 pandemic in Europe. J Eur Public Policy. 2022;29(3):321–41.
22. von Hammerstein AL, Aebi C, Barbey F, Berger C, Buettcher M, Casaulta C, et al. Interseasonal RSV infections in Switzerland - rapid establishment of a clinician-led national reporting system (RSV EpiCH). Swiss Med Wkly. 2021 Sep;151(3536):w30057. 10.4414/SMW.2021.w30057
23. Ihira M, Yoshikawa T, Ohashi M, Enomono Y, Akimoto S, Suga S, et al. Variation of human herpesvirus 7 shedding in saliva. J Infect Dis. 2003 Nov;188(9):1352–4.
24. Lewandowska DW, Schreiber PW, Schuurmans MM, Ruehe B, Zagordi O, Bayard C, et al. Metagenomic sequencing complements routine diagnostics in identifying viral pathogens in lung transplant recipients with unknown etiology of respiratory infection. ["Schildgen, Oliver"], editors. PLoS ONE. 2017;12: e0177340. doi:
25. Charalampous T, Kay GL, Richardson H, Aydin A, Baldan R, Jeanes C, et al. Nanopore metagenomics enables rapid clinical diagnosis of bacterial lower respiratory infection. Nat Biotechnol. 2019 Jul;37(7):783–92.
26. Pichler I, Schmutz S, Ziltener G, Zaheri M, Kufner V, Trkola A, et al. Rapid and sensitive single-sample viral metagenomics using Nanopore Flongle sequencing. J Virol Methods. 2023 Oct;320:114784.
The appendix is available in the pdf version of the article at https://doi.org/10.57187/s.3797.